-
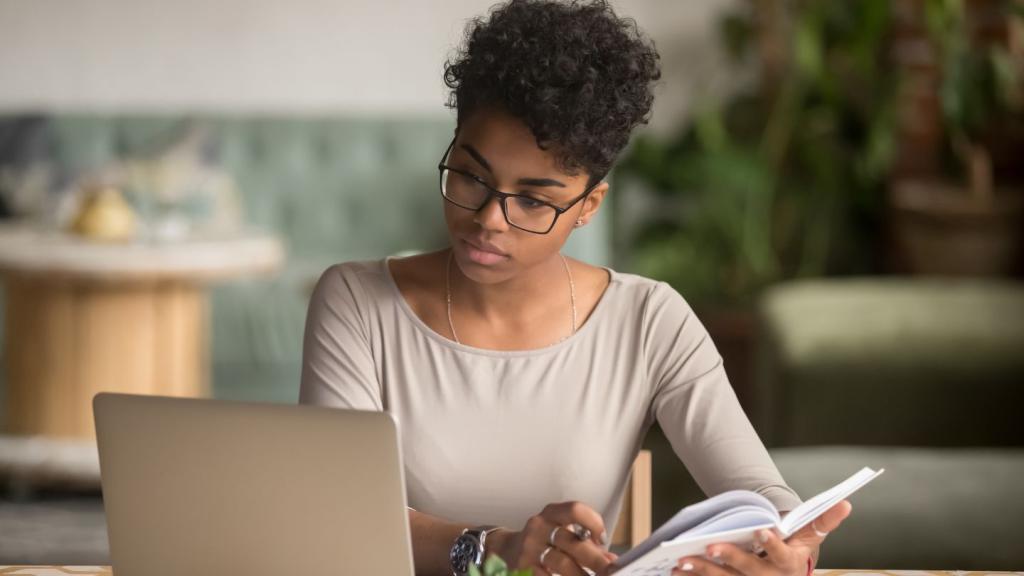
Understanding primary immunodeficiency (PI)

Understanding PI
The more you understand about primary immunodeficiency (PI), the better you can manage it. Learn about PI diagnoses and treatment options.
-
Living with PI
-
Explaining your diagnosis
- Nurturing your health
- Get support
- For parents and guardians
- Navigating insurance
-
Thriving at work
-
Traveling safely
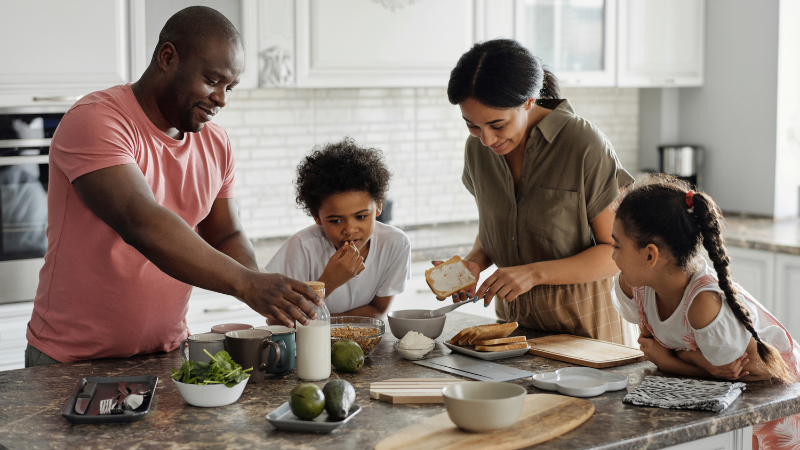

Living with PI
Living with primary immunodeficiency (PI) can be challenging, but you’re not alone—many people with PI lead full and active lives. With the right support and resources, you can, too.
-
Explaining your diagnosis
-
Get involved
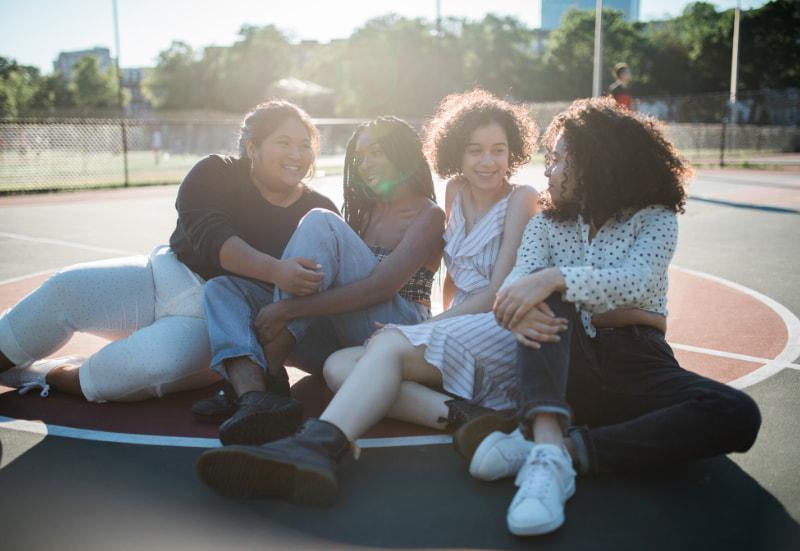

Get involved
Be a hero for those with PI. Change lives by promoting primary immunodeficiency (PI) awareness and taking action in your community through advocacy, donating, volunteering, or fundraising.
-
Advancing research and clinical care
-
Research Grant Program
-
Consulting immunologist
-
Diagnosing PI
-
Getting prior authorization
-
Clinician education
-
Survey research
-
Participating in clinical trials
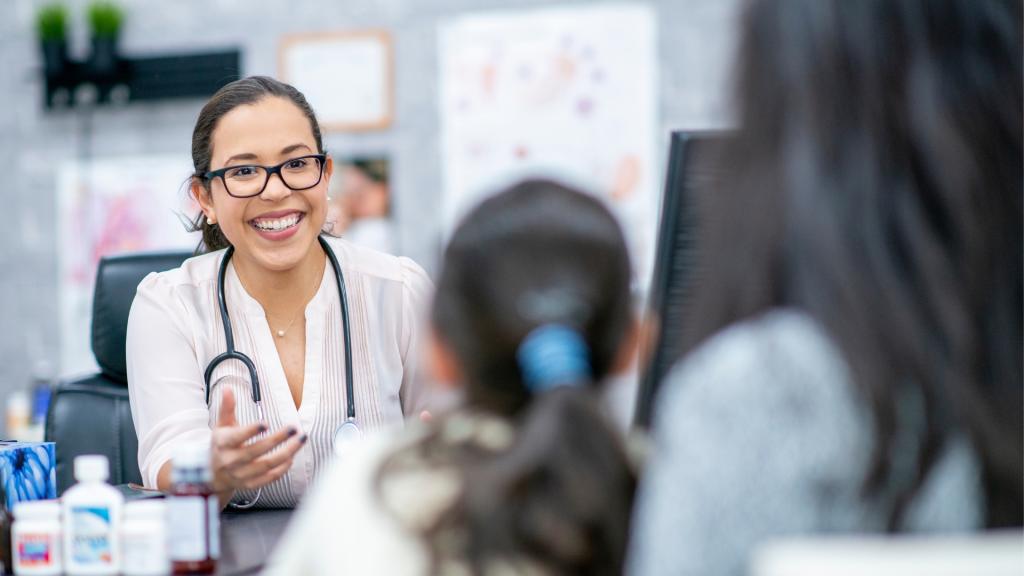

Advancing research and clinical care
Whether you’re a clinician, researcher, or an individual with primary immunodeficiency (PI), IDF has resources to help you advance the field. Get details on surveys, grants, and clinical trials.
-
Research Grant Program
For 2023, the IDF Research Grant Program is awarding more than $190,000 in support of five projects that advance the diagnosis, knowledge of underlying biology, and potential treatments for primary immunodeficiencies (PI). This year's Michael Blaese Research Grant Award, given to the top-scoring applicant, goes to Dr. Donald Kohn, distinguished professor at the University of California, Los Angeles (UCLA), and director of the UCLA David Geffen School of Medicine Human Gene and Cell Therapy Program. Read about his work on gene editing to treat X-linked agammaglobulinemia (XLA) in this companion article.
Understanding T cells in Schimke immune-osseous dysplasia (SIOD)
Awardee Dr. Benjamin Solomon, who is a fellow in allergy and immunology at Stanford University, is working on the other end of the research spectrum, trying to better understand a rare, autosomal recessive combined immunodeficiency called Schimke immune-osseous dysplasia (SIOD). Individuals with SIOD have poor bone growth, kidney function that gets worse over time, and an increased risk of severe and recurring infections. Many of these individuals die before the age of 10.
Those with SIOD are unable to fight infection because their T cells, a type of white blood cell, don’t work properly. However, those with SIOD do have circulating T cells in their blood. Solomon plans to look at a protein on the circulating T cells’ surfaces, called the T cell receptor (TCR), that he suspects differs in those with SOID and may explain their symptoms.
“Many of the most impactful treatments for primary immune deficiency come from understanding the underlying biology of those conditions,” explained Solomon.
In people without PI, each T cell makes one version of TCR, and that version determines what protein or other biological molecule the T cell will recognize. The diversity of TCRs across all of a person’s T cells is called their TCR repertoire and allows their immune system to recognize many thousands of different molecules.
Thymocytes, which ultimately develop into T cells, move from the blood marrow to the thymus to undergo two important TCR quality control processes before they are released to circulate in the blood. The first tests whether the thymocyte’s TCR is capable of recognizing a foreign molecule, which ensures that released T cells will actually protect against infection. The second process weeds out thymocytes whose TCR inappropriately recognizes ‘self’ molecules—that is, those that could trigger autoimmune symptoms.
Solomon plans to compare the diversity of TCRs found in SIOD circulating T cells with public datasets from unaffected individuals that represent different stages in T cell development. He predicts that the SIOD TCR repertoire will most closely match that of thymocytes in a person without PI before they have been through the quality control steps and released from the thymus. If Solomon shows that the TCR repertoires of circulating T cells from individuals with SIOD are most similar to thymocytes from individuals without PI, this may explain both the susceptibility to infection and the autoimmunity seen in individuals with SIOD. In essence, the immune issues of those with SOID may be the result of a lack of T cell quality control.
In addition, the techniques Solomon is using overcome a significant barrier in researchers’ ability to compare TCR repertoires in general. “This IDF award supports work to develop new statistical and computational methods capable of revealing the biology of even the rarest immune deficiencies, leading to new avenues of diagnosis and treatment,” he said.
Looking for structural and copy number variants in the DNA of individuals without a genetic diagnosis
Dr. Xiao Peng, assistant professor at Johns Hopkins University School of Medicine, is interested in finding answers for individuals with suspected PI that do not have a genetic diagnosis. Even using whole exome sequencing, which captures the sequence of all protein-coding regions of a person’s DNA, PI-causing genetic variants are found for only about 30% of those with suspected PI. Said Peng, “Too many of our patients remain baffling cases carrying many clinical labels and yet not the right ones to help get them to the diagnosis and management needed.”
One problem is that the most commonly used technique for sequencing DNA, short-read sequencing (SRS), has limitations. It can detect only small variants—single letter changes (known as single nucleotide polymorphisms), or short insertions or deletions in a stretch of DNA. In addition, raw SRS data has to be assembled like a jigsaw puzzle. Some regions of human DNA are so similar to each other that they are impossible to accurately assemble from SRS data alone.
To overcome these limitations and find additional PI-causing genetic variants, Peng plans to use specialized technology called optical genetic mapping (OGM) followed by targeted long-read sequencing (LRS). OGM and LRS will help her look for duplications of large regions of DNA, large insertions and deletions, and places where a section of DNA has been flipped around. They will also allow her to look at regions of human DNA that can’t usually be analyzed with SRS data. Peng predicted that her approach “will help more undiagnosed patients receive research-based sequencing towards the goal of getting them to molecular diagnosis.”
Peng also wants to compare OGM and LRS to more commonly used techniques to determine each technique’s ‘blind spots.’ This information will help PI researchers in general understand which technique is best and most cost-effective for detecting each type of genetic variant. Ultimately, this analysis may result in a standardized clinical approach to genetic diagnosis, with first-tier and second-tier techniques that capture a wider variety of genetic variants and provide more individuals with the genetic basis for their symptoms.
Tracking gut microbiome changes in individuals with CGD who undergo fecal microbiome transplant
Dr. Brynn O’Laughlin is a fellow in pediatric gastroenterology at Children’s National Hospital. She is using her IDF grant to track changes in the microorganism communities in a person’s digestive tract (known as the gut microbiome) before and after fecal microbiome transplant (FMT) to treat chronic granulomatous disease-associated colitis (CGD-AC). CGD-AC is an inflammatory bowel disease-like complication that occurs in approximately 50% of people with CGD. Researchers suspect that CGD-AC results from imbalances in the gut microbiome, including the types and amounts of different microorganisms and the substances they produce.
As part of an ongoing clinical trial, 20 individuals with CGD-AC will receive an FMT as a potential treatment. In FMT, the bacteria and other microorganisms from a healthy person’s digestive tract are transferred to an individual who has an unhealthy gut microbiome. The goal is to replace the unhealthy microbial community with the healthy one.
However, the clinical trial design focuses on markers of inflammation and symptom improvement, and does not include detailed analyses of how the composition of the gut microbiome changes, or not, in participants. O’Laughlin plans to determine how participants’ gut microbiome changes down to the species level, as well as want kinds of substances the microbiome is producing as it changes. These complementary analyses “will allow for the assessment of correlations between microbiome/metabolome changes and clinical outcomes after FMT,” O’Laughlin said.
In the long term, O’Laughlin’s research may help ”in the design of live biotherapeutic products for the treatment of CGD-AC if the effective components of FMT are uncovered.” Such a product, which might consist of a specific, precise mix of microorganisms, could be more reliably manufactured than the preparations that are currently used in FMT.
Identifying T cell markers that correlate with clinical severity of X-linked hyper IgM syndrome
Dr. Junghee Jenny Shin, assistant professor at Yale University, wants to be able to predict the severity of clinical symptoms for individuals with CD40 ligand deficiency (CD40LD), also known as X-linked hyper IgM syndrome, based on the properties of their T cells. Individuals with CD40LD have variants that affect the production or function of the CD40L protein on the surface of T cells. Typically, CD40L binds to a receptor on B cells, telling them to switch from making the IgM type of antibodies to IgA, IgE, or IgG. Because this switch doesn’t happen without functioning CD40L, those with CD40LD have high IgM levels, but low levels of other types of antibodies.
Symptoms of CD40LD include severe infections starting in the first year or two of life, as well as an increased risk of cancer. While the median age of survival is 25, there is a wide range in clinical severity of the disease. Shin wants to link the genetic variant each individual with CD40LD has with their disease severity and data showing how their T cells differ from those of unaffected individuals.
To do this, Shin plans to look at the genes turned on and off and the proteins made in several types of T cells in individuals with CD40LD compared to those who are unaffected. This broad view of how T cells are affected will help identify “distinct T cell characteristics and function in CD40LD in relation to [patients’] comorbidities, [which] will facilitate the development of precisely defined treatment strategies that can limit further infection, prevent the development of malignancy, and improve survival.”
Related resources
Sign up for updates from IDF
Receive news and helpful resources to your cell phone or inbox. You can change or cancel your subscription at any time.




The Immune Deficiency Foundation improves the diagnosis, treatment, and quality of life for every person affected by primary immunodeficiency.
We foster a community that is connected, engaged, and empowered through advocacy, education, and research.